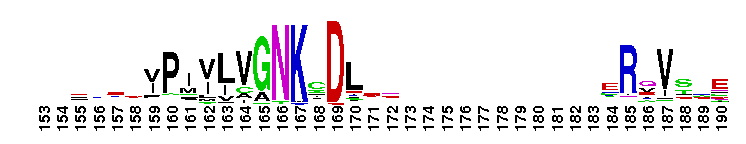| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
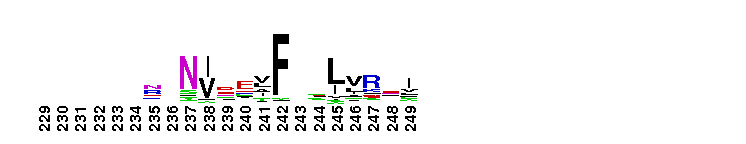
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




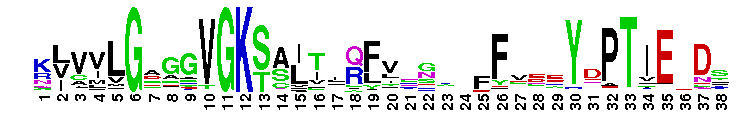
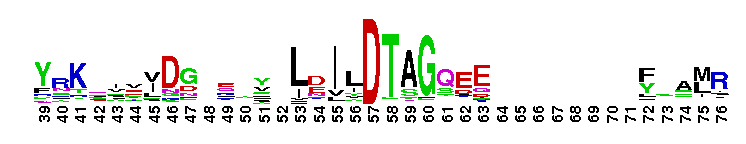
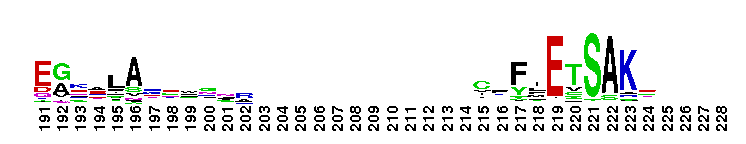
 Rat sarcoma (Ras) family of small guanosine triphosphatases (GTPases). The Ras family of the Ras superfamily includes classical N-Ras, H-Ras, and K-Ras, as well as R-Ras, Rap, Ral, Rheb, Rhes, ARHI, RERG, Rin/Rit, RSR1, RRP22, Ras2, Ras-dva, and RGK proteins. Ras proteins regulate cell growth, proliferation and differentiation. Ras is activated by guanine nucleotide exchange factors (GEFs) that release GDP and allow GTP binding. Many RasGEFs have been identified. These are sequestered in the cytosol until activation by growth factors triggers recruitment to the plasma membrane or Golgi, where the GEF colocalizes with Ras. Active GTP-bound Ras interacts with several effector proteins: among the best characterized are the Raf kinases, phosphatidylinositol 3-kinase (PI3K), RalGEFs and NORE/MST1. Most Ras proteins contain a lipid modification site at the C-terminus, with a typical sequence motif CaaX, where a = an aliphatic amino acid and X = any amino acid. Lipid binding is essential for membrane attachment, a key feature of most Ras proteins. Due to the presence of truncated sequences in this CD, the lipid modification site is not available for annotation.
Rat sarcoma (Ras) family of small guanosine triphosphatases (GTPases). The Ras family of the Ras superfamily includes classical N-Ras, H-Ras, and K-Ras, as well as R-Ras, Rap, Ral, Rheb, Rhes, ARHI, RERG, Rin/Rit, RSR1, RRP22, Ras2, Ras-dva, and RGK proteins. Ras proteins regulate cell growth, proliferation and differentiation. Ras is activated by guanine nucleotide exchange factors (GEFs) that release GDP and allow GTP binding. Many RasGEFs have been identified. These are sequestered in the cytosol until activation by growth factors triggers recruitment to the plasma membrane or Golgi, where the GEF colocalizes with Ras. Active GTP-bound Ras interacts with several effector proteins: among the best characterized are the Raf kinases, phosphatidylinositol 3-kinase (PI3K), RalGEFs and NORE/MST1. Most Ras proteins contain a lipid modification site at the C-terminus, with a typical sequence motif CaaX, where a = an aliphatic amino acid and X = any amino acid. Lipid binding is essential for membrane attachment, a key feature of most Ras proteins. Due to the presence of truncated sequences in this CD, the lipid modification site is not available for annotation. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.