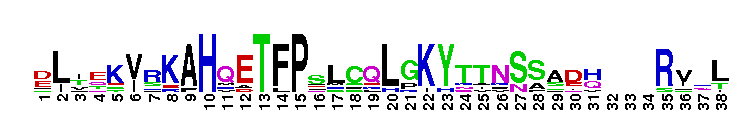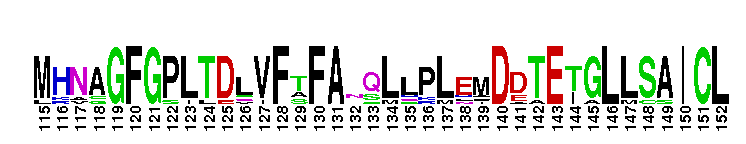| ||||||||||||||||||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




 The ligand binding domain (LBD) of retinoic acid receptor (RAR), a members of the nuclear receptor superfamily. The ligand binding domain (LBD) of retinoic acid receptor (RAR): Retinoic acid receptors are members of the nuclear receptor (NR) superfamily of ligand-regulated transcription factors. RARs mediate the biological effect of retinoids, including both naturally dietary vitamin A (retinol) metabolites and active synthetic analogs. Retinoids play key roles in a wide variety of essential biological processes, such as vertebrate embryonic morphogenesis and organogenesis, differentiation and apoptosis, and homeostasis. RARs function as heterodimers with retinoic X receptors by binding to specific RAR response elements (RAREs) found in the promoter regions of retinoid target genes. In the absence of ligand, the RAR-RXR heterodimer recruits the corepressor proteins NCoR or AMRT, and associated factors such as histone deacetylases or DNA-methyltransferases, leading to an inactive condensed chromatin structure, preventing transcription. Upon ligand binding, the corepressors are released, and coactivator complexes such as histone acetyltransferase or histone arginine methyltransferases are recruited to activate transcription. There are three RAR subtypes (alpha, beta, gamma), originating from three distinct genes. For each subtype, several isoforms exist that differ in their N-terminal region, allowing retinoids to exert their pleiotropic effects. Like other members of the nuclear receptor (NR) superfamily of ligand-activated transcription factors, retinoic acid receptors have a central well conserved DNA binding domain (DBD), a variable N-terminal domain, a non-conserved hinge and a C-terminal ligand binding domain (LBD).
The ligand binding domain (LBD) of retinoic acid receptor (RAR), a members of the nuclear receptor superfamily. The ligand binding domain (LBD) of retinoic acid receptor (RAR): Retinoic acid receptors are members of the nuclear receptor (NR) superfamily of ligand-regulated transcription factors. RARs mediate the biological effect of retinoids, including both naturally dietary vitamin A (retinol) metabolites and active synthetic analogs. Retinoids play key roles in a wide variety of essential biological processes, such as vertebrate embryonic morphogenesis and organogenesis, differentiation and apoptosis, and homeostasis. RARs function as heterodimers with retinoic X receptors by binding to specific RAR response elements (RAREs) found in the promoter regions of retinoid target genes. In the absence of ligand, the RAR-RXR heterodimer recruits the corepressor proteins NCoR or AMRT, and associated factors such as histone deacetylases or DNA-methyltransferases, leading to an inactive condensed chromatin structure, preventing transcription. Upon ligand binding, the corepressors are released, and coactivator complexes such as histone acetyltransferase or histone arginine methyltransferases are recruited to activate transcription. There are three RAR subtypes (alpha, beta, gamma), originating from three distinct genes. For each subtype, several isoforms exist that differ in their N-terminal region, allowing retinoids to exert their pleiotropic effects. Like other members of the nuclear receptor (NR) superfamily of ligand-activated transcription factors, retinoic acid receptors have a central well conserved DNA binding domain (DBD), a variable N-terminal domain, a non-conserved hinge and a C-terminal ligand binding domain (LBD). No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.