| |||||||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




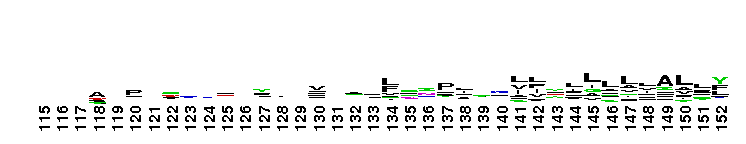
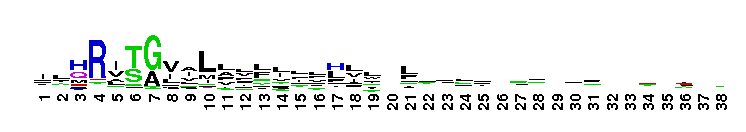
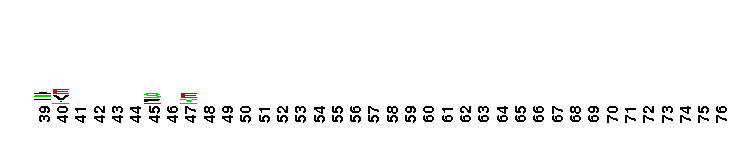
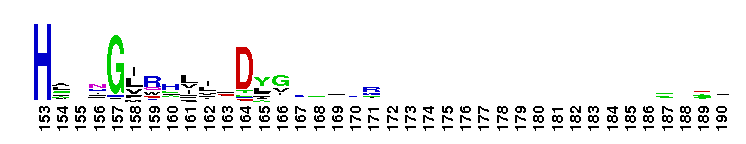
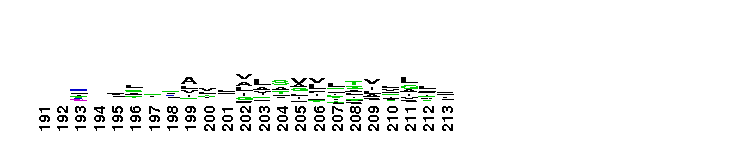
 Succinate:quinone oxidoreductase (SQR) and Quinol:fumarate reductase (QFR) family, transmembrane subunits; SQR catalyzes the oxidation of succinate to fumarate coupled to the reduction of quinone to quinol, while QFR catalyzes the reverse reaction. SQR, also called succinate dehydrogenase or Complex II, is part of the citric acid cycle and the aerobic respiratory chain, while QFR is involved in anaerobic respiration with fumarate as the terminal electron acceptor. SQRs may reduce either high or low potential quinones while QFRs oxidize only low potential quinols. SQR and QFR share a common subunit arrangement, composed of a flavoprotein catalytic subunit, an iron-sulfur protein and one or two hydrophobic transmembrane subunits. The structural arrangement allows efficient electron transfer between the catalytic subunit, through iron-sulfur centers, and the transmembrane subunit(s) containing the electron donor/acceptor (quinol or quinone). The reversible reduction of quinone is an essential feature of respiration, allowing the transfer of electrons between respiratory complexes. SQRs and QFRs can be classified into five types (A-E) according to the number of their hydrophobic subunits and heme groups. This classification is consistent with the characteristics and phylogeny of the catalytic and iron-sulfur subunits. Type E proteins, e.g. non-classical archael SQRs, contain atypical transmembrane subunits and are not included in this hierarchy. The heme and quinone binding sites reside in the transmembrane subunits. Although succinate oxidation and fumarate reduction are carried out by separate enzymes in most organisms, some bifunctional enzymes that exhibit both SQR and QFR activities exist.
Succinate:quinone oxidoreductase (SQR) and Quinol:fumarate reductase (QFR) family, transmembrane subunits; SQR catalyzes the oxidation of succinate to fumarate coupled to the reduction of quinone to quinol, while QFR catalyzes the reverse reaction. SQR, also called succinate dehydrogenase or Complex II, is part of the citric acid cycle and the aerobic respiratory chain, while QFR is involved in anaerobic respiration with fumarate as the terminal electron acceptor. SQRs may reduce either high or low potential quinones while QFRs oxidize only low potential quinols. SQR and QFR share a common subunit arrangement, composed of a flavoprotein catalytic subunit, an iron-sulfur protein and one or two hydrophobic transmembrane subunits. The structural arrangement allows efficient electron transfer between the catalytic subunit, through iron-sulfur centers, and the transmembrane subunit(s) containing the electron donor/acceptor (quinol or quinone). The reversible reduction of quinone is an essential feature of respiration, allowing the transfer of electrons between respiratory complexes. SQRs and QFRs can be classified into five types (A-E) according to the number of their hydrophobic subunits and heme groups. This classification is consistent with the characteristics and phylogeny of the catalytic and iron-sulfur subunits. Type E proteins, e.g. non-classical archael SQRs, contain atypical transmembrane subunits and are not included in this hierarchy. The heme and quinone binding sites reside in the transmembrane subunits. Although succinate oxidation and fumarate reduction are carried out by separate enzymes in most organisms, some bifunctional enzymes that exhibit both SQR and QFR activities exist. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.