| ||||||||||||||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




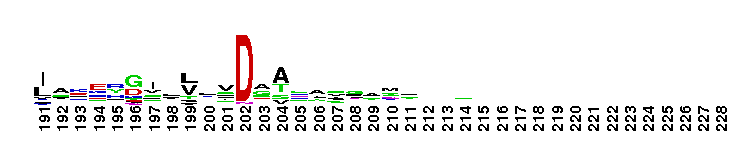
 Aspartate aminotransferase (AAT) superfamily (fold type I) of pyridoxal phosphate (PLP)-dependent enzymes. PLP combines with an alpha-amino acid to form a compound called a Schiff base or aldimine intermediate, which depending on the reaction, is the substrate in four kinds of reactions (1) transamination (movement of amino groups), (2) racemization (redistribution of enantiomers), (3) decarboxylation (removing COOH groups), and (4) various side-chain reactions depending on the enzyme involved. Pyridoxal phosphate (PLP) dependent enzymes were previously classified into alpha, beta and gamma classes, based on the chemical characteristics (carbon atom involved) of the reaction they catalyzed. The availability of several structures allowed a comprehensive analysis of the evolutionary classification of PLP dependent enzymes, and it was found that the functional classification did not always agree with the evolutionary history of these enzymes. Structure and sequence analysis has revealed that the PLP dependent enzymes can be classified into four major groups of different evolutionary origin: aspartate aminotransferase superfamily (fold type I), tryptophan synthase beta superfamily (fold type II), alanine racemase superfamily (fold type III), and D-amino acid superfamily (fold type IV) and Glycogen phophorylase family (fold type V).
Aspartate aminotransferase (AAT) superfamily (fold type I) of pyridoxal phosphate (PLP)-dependent enzymes. PLP combines with an alpha-amino acid to form a compound called a Schiff base or aldimine intermediate, which depending on the reaction, is the substrate in four kinds of reactions (1) transamination (movement of amino groups), (2) racemization (redistribution of enantiomers), (3) decarboxylation (removing COOH groups), and (4) various side-chain reactions depending on the enzyme involved. Pyridoxal phosphate (PLP) dependent enzymes were previously classified into alpha, beta and gamma classes, based on the chemical characteristics (carbon atom involved) of the reaction they catalyzed. The availability of several structures allowed a comprehensive analysis of the evolutionary classification of PLP dependent enzymes, and it was found that the functional classification did not always agree with the evolutionary history of these enzymes. Structure and sequence analysis has revealed that the PLP dependent enzymes can be classified into four major groups of different evolutionary origin: aspartate aminotransferase superfamily (fold type I), tryptophan synthase beta superfamily (fold type II), alanine racemase superfamily (fold type III), and D-amino acid superfamily (fold type IV) and Glycogen phophorylase family (fold type V). No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.