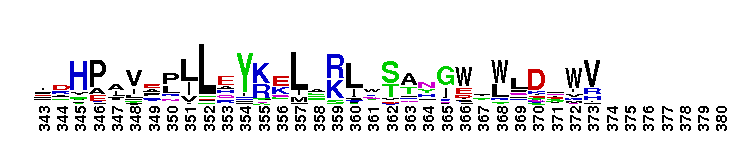| |||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




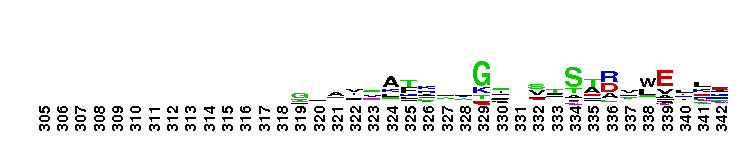
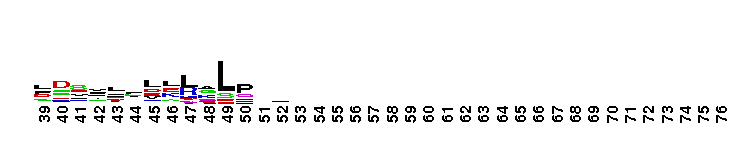
 Family A polymerase primarily fills DNA gaps that arise during DNA repair, recombination and replication. DNA polymerase family A, 5'-3' polymerase domain. Family A polymerase functions primarily to fill DNA gaps that arise during DNA repair, recombination and replication. DNA-dependent DNA polymerases can be classified into six main groups based upon phylogenetic relationships with E. coli polymerase I (classA), E. coli polymerase II (class B), E.coli polymerase III (class C), euryarchaeota polymerase II (class D), human polymerase beta (class X), E. coli UmuC/DinB and eukaryotic RAP 30/Xeroderma pigmentosum variant (class Y). Family A polymerases are found primarily in organisms related to prokaryotes and include prokaryotic DNA polymerase I, mitochondrial polymerase gamma, and several bacteriophage polymerases including those from odd-numbered phage (T3, T5, and T7). Prokaryotic polymerase I (pol I) has two functional domains located on the same polypeptide; a 5'-3' polymerase and a 5'-3' exonuclease. Pol I uses its 5' nuclease activity to remove the ribonucleotide portion of newly synthesized Okazaki fragments and the DNA polymerase activity to fill in the resulting gap. The structure of these polymerases resembles in overall morphology a cupped human right hand, with fingers (which bind an incoming nucleotide and interact with the single-stranded template), palm (which harbors the catalytic amino acid residues and also binds an incoming dNTP) and thumb (which binds double-stranded DNA) subdomains.
Family A polymerase primarily fills DNA gaps that arise during DNA repair, recombination and replication. DNA polymerase family A, 5'-3' polymerase domain. Family A polymerase functions primarily to fill DNA gaps that arise during DNA repair, recombination and replication. DNA-dependent DNA polymerases can be classified into six main groups based upon phylogenetic relationships with E. coli polymerase I (classA), E. coli polymerase II (class B), E.coli polymerase III (class C), euryarchaeota polymerase II (class D), human polymerase beta (class X), E. coli UmuC/DinB and eukaryotic RAP 30/Xeroderma pigmentosum variant (class Y). Family A polymerases are found primarily in organisms related to prokaryotes and include prokaryotic DNA polymerase I, mitochondrial polymerase gamma, and several bacteriophage polymerases including those from odd-numbered phage (T3, T5, and T7). Prokaryotic polymerase I (pol I) has two functional domains located on the same polypeptide; a 5'-3' polymerase and a 5'-3' exonuclease. Pol I uses its 5' nuclease activity to remove the ribonucleotide portion of newly synthesized Okazaki fragments and the DNA polymerase activity to fill in the resulting gap. The structure of these polymerases resembles in overall morphology a cupped human right hand, with fingers (which bind an incoming nucleotide and interact with the single-stranded template), palm (which harbors the catalytic amino acid residues and also binds an incoming dNTP) and thumb (which binds double-stranded DNA) subdomains. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.