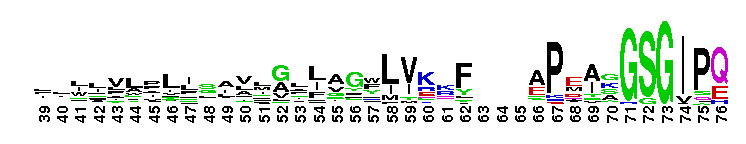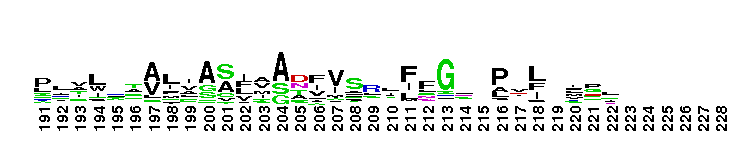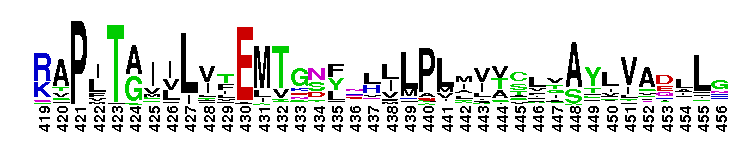| ||||||||||||||||||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




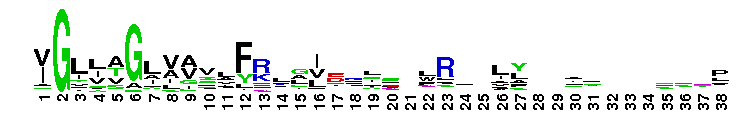
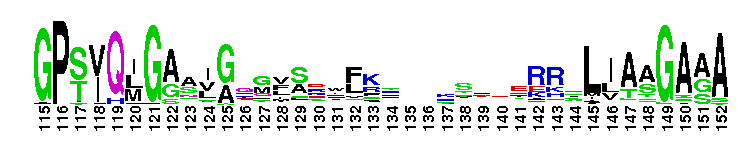
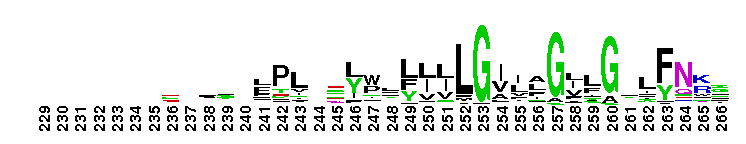
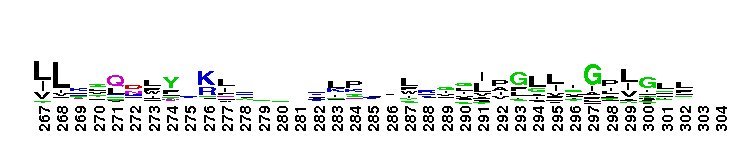
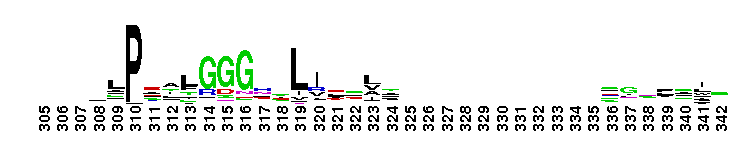
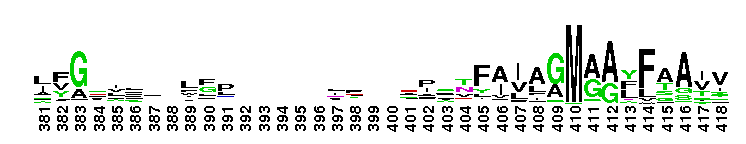
 ClC chloride channel EriC. This domain is found in the EriC chloride transporters that mediate the extreme acid resistance response in eubacteria and archaea. This response allows bacteria to survive in the acidic environments by decarboxylation-linked proton utilization. As shown for Escherichia coli EriC, these channels can counterbalance the electric current produced by the outwardly directed virtual proton pump linked to amino acid decarboxylation. The EriC proteins belong to the ClC superfamily of chloride ion channels, which share a unique double-barreled architecture and voltage-dependent gating mechanism. The voltage-dependent gating is conferred by the permeating anion itself, acting as the gating charge. In Escherichia coli EriC, a glutamate residue that protrudes into the pore is thought to participate in gating by binding to a Cl- ion site within the selectivity filter.
ClC chloride channel EriC. This domain is found in the EriC chloride transporters that mediate the extreme acid resistance response in eubacteria and archaea. This response allows bacteria to survive in the acidic environments by decarboxylation-linked proton utilization. As shown for Escherichia coli EriC, these channels can counterbalance the electric current produced by the outwardly directed virtual proton pump linked to amino acid decarboxylation. The EriC proteins belong to the ClC superfamily of chloride ion channels, which share a unique double-barreled architecture and voltage-dependent gating mechanism. The voltage-dependent gating is conferred by the permeating anion itself, acting as the gating charge. In Escherichia coli EriC, a glutamate residue that protrudes into the pore is thought to participate in gating by binding to a Cl- ion site within the selectivity filter. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.