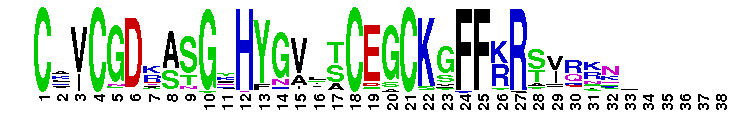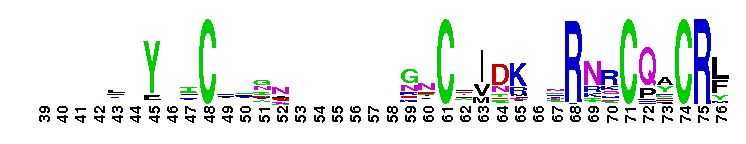| |||||||||||||||||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




 DNA-binding domain of nuclear receptors is composed of two C4-type zinc fingers. DNA-binding domain of nuclear receptors is composed of two C4-type zinc fingers. Each zinc finger contains a group of four Cys residues which co-ordinates a single zinc atom. It interacts with a specific DNA site upstream of the target gene and modulates the rate of transcriptional initiation. Nuclear receptors form a superfamily of ligand-activated transcription regulators, which regulate various physiological functions, from development, reproduction, to homeostasis and metabolism in animals (metazoans). The family contains not only receptors for known ligands but also orphan receptors for which ligands do not exist or have not been identified. NRs share a common structural organization with a central well conserved DNA binding domain (DBD), a variable N-terminal domain, a flexible hinge and a C-terminal ligand binding domain (LBD). Most nuclear receptors bind as homodimers or heterodimers to their target sites, which consist of two hexameric half-sites. Specificity is determined by the half-site sequence, the relative orientation of the half-sites and the number of spacer nucleotides between the half-sites. However, a growing number of nuclear receptors have been reported to bind to DNA as monomers.
DNA-binding domain of nuclear receptors is composed of two C4-type zinc fingers. DNA-binding domain of nuclear receptors is composed of two C4-type zinc fingers. Each zinc finger contains a group of four Cys residues which co-ordinates a single zinc atom. It interacts with a specific DNA site upstream of the target gene and modulates the rate of transcriptional initiation. Nuclear receptors form a superfamily of ligand-activated transcription regulators, which regulate various physiological functions, from development, reproduction, to homeostasis and metabolism in animals (metazoans). The family contains not only receptors for known ligands but also orphan receptors for which ligands do not exist or have not been identified. NRs share a common structural organization with a central well conserved DNA binding domain (DBD), a variable N-terminal domain, a flexible hinge and a C-terminal ligand binding domain (LBD). Most nuclear receptors bind as homodimers or heterodimers to their target sites, which consist of two hexameric half-sites. Specificity is determined by the half-site sequence, the relative orientation of the half-sites and the number of spacer nucleotides between the half-sites. However, a growing number of nuclear receptors have been reported to bind to DNA as monomers. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.