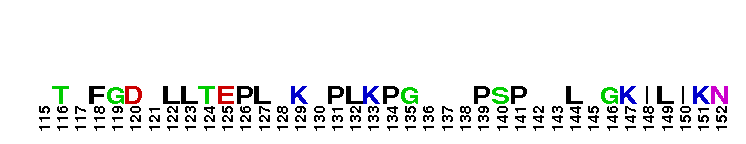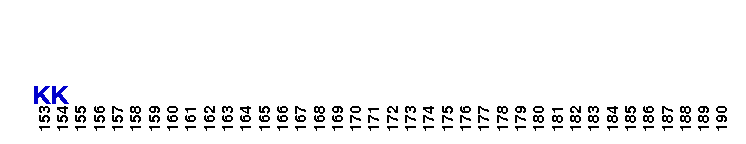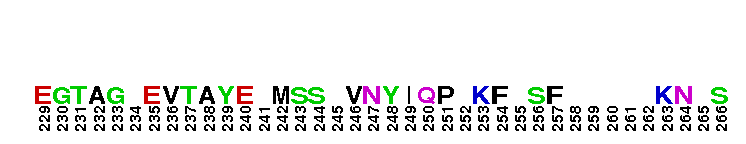| ||||||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




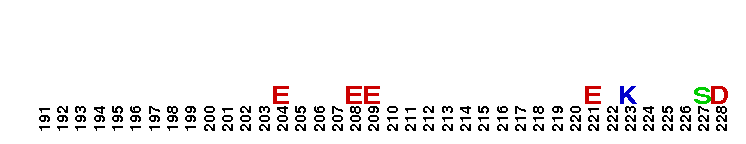
 Catalytic domain of metazoan phosphoinositide-specific phospholipase C-beta2. This subfamily corresponds to the catalytic domain present in metazoan phosphoinositide-specific phospholipase C (PI-PLC, EC 3.1.4.11)-beta isozyme 2. PI-PLC is a signaling enzyme that hydrolyzes the membrane phospholipids phosphatidylinositol-4,5-bisphosphate (PIP2) to generate two important second messengers in eukaryotic signal transduction cascades, Inositol 1,4,5-trisphosphate (InsP3) and diacylglycerol (DAG). InsP3 triggers inflow of calcium from intracellular stores, while DAG, together with calcium, activates protein kinase C, which goes on to phosphorylate other molecules, leading to altered cellular activity. Calcium is required for the catalysis. PLC-beta represents a class of mammalian PI-PLC that has an N-terminal pleckstrin homology (PH) domain, an array of EF hands, a PLC catalytic core domain, a C2 domain, and a unique C-terminal coiled-coil (CT) domain necessary for homodimerization. The PLC catalytic core domain is a TIM barrel with two highly conserved regions (X and Y) split by a highly degenerate linker sequence. PI-PLC-beta2 is expressed at highest levels in cells of hematopoietic origin. It is activated by the heterotrimeric G protein alpha q subunits through their C2 domain and long C-terminal extension. It is also activated by the beta-gamma subunits of heterotrimeric G proteins.
Catalytic domain of metazoan phosphoinositide-specific phospholipase C-beta2. This subfamily corresponds to the catalytic domain present in metazoan phosphoinositide-specific phospholipase C (PI-PLC, EC 3.1.4.11)-beta isozyme 2. PI-PLC is a signaling enzyme that hydrolyzes the membrane phospholipids phosphatidylinositol-4,5-bisphosphate (PIP2) to generate two important second messengers in eukaryotic signal transduction cascades, Inositol 1,4,5-trisphosphate (InsP3) and diacylglycerol (DAG). InsP3 triggers inflow of calcium from intracellular stores, while DAG, together with calcium, activates protein kinase C, which goes on to phosphorylate other molecules, leading to altered cellular activity. Calcium is required for the catalysis. PLC-beta represents a class of mammalian PI-PLC that has an N-terminal pleckstrin homology (PH) domain, an array of EF hands, a PLC catalytic core domain, a C2 domain, and a unique C-terminal coiled-coil (CT) domain necessary for homodimerization. The PLC catalytic core domain is a TIM barrel with two highly conserved regions (X and Y) split by a highly degenerate linker sequence. PI-PLC-beta2 is expressed at highest levels in cells of hematopoietic origin. It is activated by the heterotrimeric G protein alpha q subunits through their C2 domain and long C-terminal extension. It is also activated by the beta-gamma subunits of heterotrimeric G proteins. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.