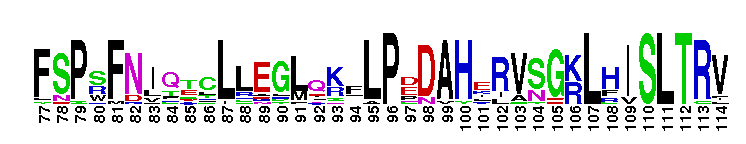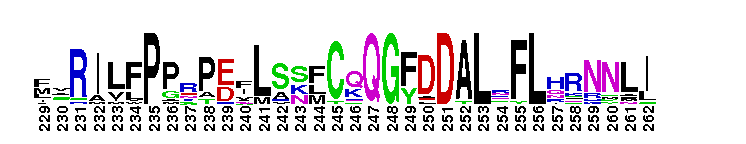| |||||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




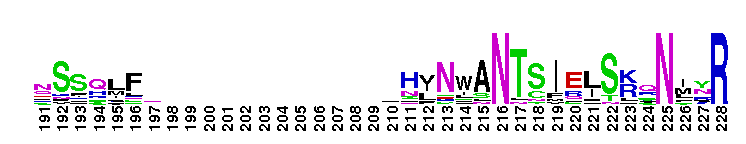
 Calcium-independent phospholipase A2; Classified as Group IVA-1 PLA2. Calcium-independent phospholipase A2; otherwise known as Group IVA-1 PLA2. It contains the lipase consensus sequence (Gly-X-Ser-X-Gly);mutagenesis experiments confirm the role of this serine as a nucleophile. Some members of this group show triacylglycerol lipase activity (EC 3:1:1:3). Members include iPLA-1, iPLA-2, and iPLA-3 from Aedes aegypti and show acylglycerol transacylase/lipase activity. Also includes putative iPLA2-eta from Pediculus humanus corporis which shows patatin-like phospholipase activity.
Calcium-independent phospholipase A2; Classified as Group IVA-1 PLA2. Calcium-independent phospholipase A2; otherwise known as Group IVA-1 PLA2. It contains the lipase consensus sequence (Gly-X-Ser-X-Gly);mutagenesis experiments confirm the role of this serine as a nucleophile. Some members of this group show triacylglycerol lipase activity (EC 3:1:1:3). Members include iPLA-1, iPLA-2, and iPLA-3 from Aedes aegypti and show acylglycerol transacylase/lipase activity. Also includes putative iPLA2-eta from Pediculus humanus corporis which shows patatin-like phospholipase activity. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.