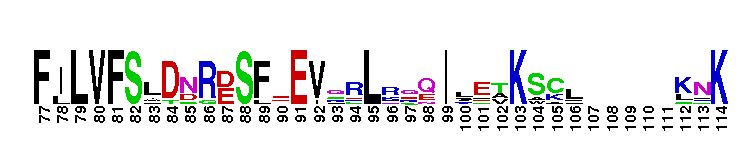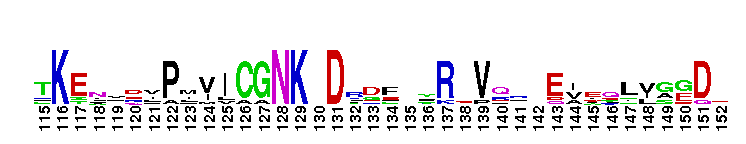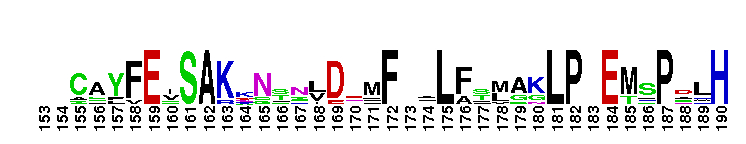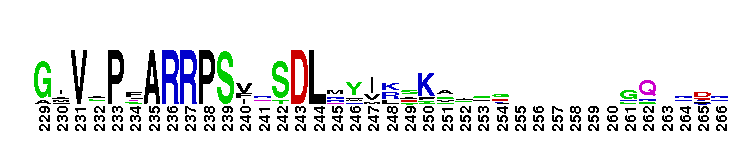| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
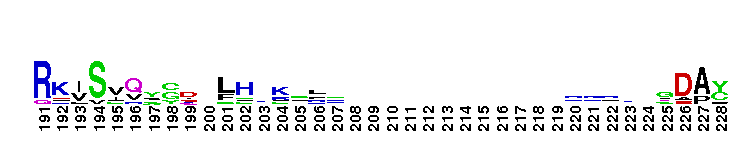
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




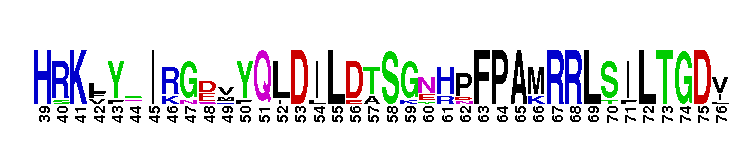
 Ras homolog enriched in striatum (Rhes) and activator of G-protein signaling 1 (Dexras1/AGS1). This subfamily includes Rhes (Ras homolog enriched in striatum) and Dexras1/AGS1 (activator of G-protein signaling 1). These proteins are homologous, but exhibit significant differences in tissue distribution and subcellular localization. Rhes is found primarily in the striatum of the brain, but is also expressed in other areas of the brain, such as the cerebral cortex, hippocampus, inferior colliculus, and cerebellum. Rhes expression is controlled by thyroid hormones. In rat PC12 cells, Rhes is farnesylated and localizes to the plasma membrane. Rhes binds and activates PI3K, and plays a role in coupling serpentine membrane receptors with heterotrimeric G-protein signaling. Rhes has recently been shown to be reduced under conditions of dopamine supersensitivity and may play a role in determining dopamine receptor sensitivity. Dexras1/AGS1 is a dexamethasone-induced Ras protein that is expressed primarily in the brain, with low expression levels in other tissues. Dexras1 localizes primarily to the cytoplasm, and is a critical regulator of the circadian master clock to photic and nonphotic input. Most Ras proteins contain a lipid modification site at the C-terminus, with a typical sequence motif CaaX, where a = an aliphatic amino acid and X = any amino acid. Lipid binding is essential for membrane attachment, a key feature of most Ras proteins.
Ras homolog enriched in striatum (Rhes) and activator of G-protein signaling 1 (Dexras1/AGS1). This subfamily includes Rhes (Ras homolog enriched in striatum) and Dexras1/AGS1 (activator of G-protein signaling 1). These proteins are homologous, but exhibit significant differences in tissue distribution and subcellular localization. Rhes is found primarily in the striatum of the brain, but is also expressed in other areas of the brain, such as the cerebral cortex, hippocampus, inferior colliculus, and cerebellum. Rhes expression is controlled by thyroid hormones. In rat PC12 cells, Rhes is farnesylated and localizes to the plasma membrane. Rhes binds and activates PI3K, and plays a role in coupling serpentine membrane receptors with heterotrimeric G-protein signaling. Rhes has recently been shown to be reduced under conditions of dopamine supersensitivity and may play a role in determining dopamine receptor sensitivity. Dexras1/AGS1 is a dexamethasone-induced Ras protein that is expressed primarily in the brain, with low expression levels in other tissues. Dexras1 localizes primarily to the cytoplasm, and is a critical regulator of the circadian master clock to photic and nonphotic input. Most Ras proteins contain a lipid modification site at the C-terminus, with a typical sequence motif CaaX, where a = an aliphatic amino acid and X = any amino acid. Lipid binding is essential for membrane attachment, a key feature of most Ras proteins. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.