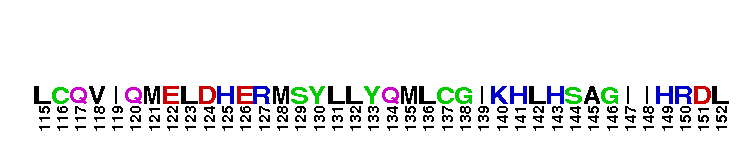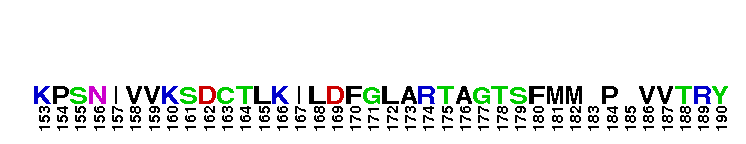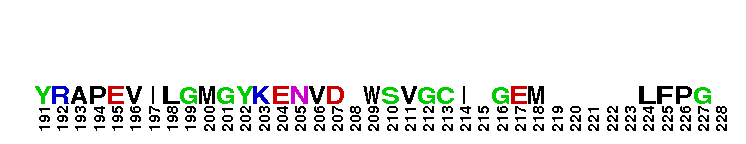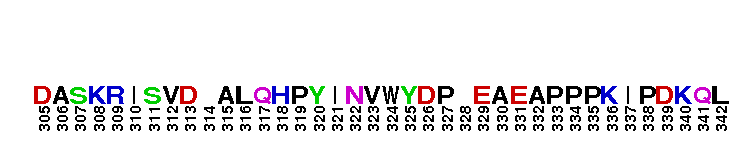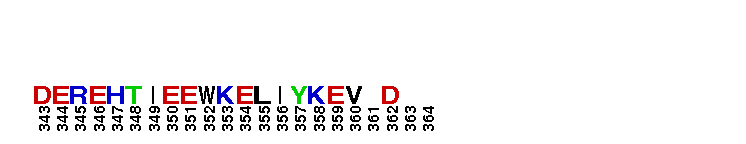| |||||||||||||||||||||||||||||||||||||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




 Catalytic domain of the Serine/Threonine Kinase, c-Jun N-terminal Kinase 1. Serine/Threonine Kinases (STKs), c-Jun N-terminal kinase 1 (JNK1) subfamily, catalytic (c) domain. STKs catalyze the transfer of the gamma-phosphoryl group from ATP to serine/threonine residues on protein substrates. The JNK1 subfamily is part of a larger superfamily that includes the catalytic domains of other protein STKs, protein tyrosine kinases, RIO kinases, aminoglycoside phosphotransferase, choline kinase, and phosphoinositide 3-kinase. JNKs are mitogen-activated protein kinases (MAPKs) that are involved in many stress-activated responses including those during inflammation, neurodegeneration, apoptosis, and persistent pain sensitization, among others. Vetebrates harbor three different JNK genes (Jnk1, Jnk2, and Jnk3). JNK1, like JNK2, is expressed in every cell and tissue type. Initially it was thought that JNK1 and JNK2 were functionally redundant as mice deficient in either genes (Jnk1 or Jnk2) could survive but disruption of both genes resulted in lethality. However, recent studies have shown that JNK1 and JNK2 perform distinct functions through specific binding partners and substrates. JNK1 specifically binds with JAMP (JNK1-associated membrane protein), which regulates the duration of JNK1 activity in response to stimuli. Specific JNK1 substrates include Itch and SG10, which are implicated in Th2 responses and airway inflammation, and microtubule dynamics and axodendritic length, respectively. Mice deficient in Jnk1 are protected against arthritis, obesity, type 2 diabetes, cardiac cell death, and non-alcoholic liver disease, suggesting that JNK1 may play roles in the pathogenesis of these diseases.
Catalytic domain of the Serine/Threonine Kinase, c-Jun N-terminal Kinase 1. Serine/Threonine Kinases (STKs), c-Jun N-terminal kinase 1 (JNK1) subfamily, catalytic (c) domain. STKs catalyze the transfer of the gamma-phosphoryl group from ATP to serine/threonine residues on protein substrates. The JNK1 subfamily is part of a larger superfamily that includes the catalytic domains of other protein STKs, protein tyrosine kinases, RIO kinases, aminoglycoside phosphotransferase, choline kinase, and phosphoinositide 3-kinase. JNKs are mitogen-activated protein kinases (MAPKs) that are involved in many stress-activated responses including those during inflammation, neurodegeneration, apoptosis, and persistent pain sensitization, among others. Vetebrates harbor three different JNK genes (Jnk1, Jnk2, and Jnk3). JNK1, like JNK2, is expressed in every cell and tissue type. Initially it was thought that JNK1 and JNK2 were functionally redundant as mice deficient in either genes (Jnk1 or Jnk2) could survive but disruption of both genes resulted in lethality. However, recent studies have shown that JNK1 and JNK2 perform distinct functions through specific binding partners and substrates. JNK1 specifically binds with JAMP (JNK1-associated membrane protein), which regulates the duration of JNK1 activity in response to stimuli. Specific JNK1 substrates include Itch and SG10, which are implicated in Th2 responses and airway inflammation, and microtubule dynamics and axodendritic length, respectively. Mice deficient in Jnk1 are protected against arthritis, obesity, type 2 diabetes, cardiac cell death, and non-alcoholic liver disease, suggesting that JNK1 may play roles in the pathogenesis of these diseases. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.