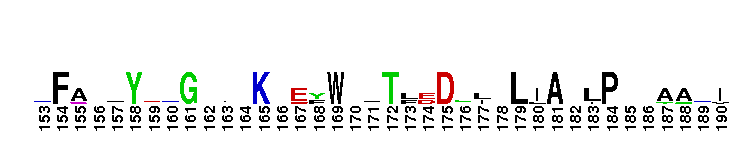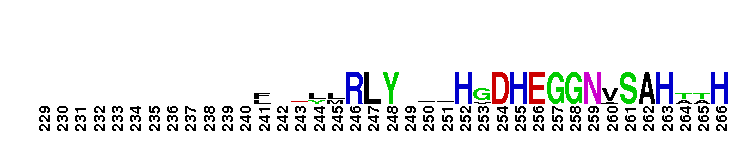| |||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




 Saccharomyces cerevisiae (Sc) 2-methylcitrate synthase Cit3-like. 2-methylcitrate synthase (2MCS) catalyzes the condensation of propionyl-coenzyme A (PrCoA) and oxaloacetate (OAA) to form 2-methylcitrate and CoA. Citrate synthase (CS) catalyzes the condensation of acetyl coenzyme A (AcCoA) with OAA to form citrate and CoA, the first step in the citric acid cycle (TCA or Krebs cycle). The overall CS reaction is thought to proceed through three partial reactions and involves both closed and open conformational forms of the enzyme: a) the carbanion or equivalent is generated from AcCoA by base abstraction of a proton, b) the nucleophilic attack of this carbanion on OAA to generate citryl-CoA, and c) the hydrolysis of citryl-CoA to produce citrate and CoA. There are two types of CSs: type I CS and type II CSs. Type I CSs are found in eukarya, gram-positive bacteria, archaea, and in some gram-negative bacteria and are homodimers with both subunits participating in the active site. Type II CSs are unique to gram-negative bacteria and are homohexamers of identical subunits (approximated as a trimer of dimers). ScCit3 is mitochondrial and functions in the metabolism of PrCoA; it is a dual specificity CS and 2MCS, having similar catalytic efficiency with both AcCoA and PrCoA. The pattern of expression of the ScCIT3 gene follows that of the major mitochondrial CS gene (CIT1, not included in this group) and its expression is increased in the presence of a CIT1 deletion. This group also contains Aspergillus nidulans 2MCS; a deletion of the gene encoding this protein results in a strain unable to grow on propionate. This group contains proteins which functions exclusively as either a CS or a 2MCS, as well as those with relaxed specificity which have dual functions as both a CS and a 2MCS.
Saccharomyces cerevisiae (Sc) 2-methylcitrate synthase Cit3-like. 2-methylcitrate synthase (2MCS) catalyzes the condensation of propionyl-coenzyme A (PrCoA) and oxaloacetate (OAA) to form 2-methylcitrate and CoA. Citrate synthase (CS) catalyzes the condensation of acetyl coenzyme A (AcCoA) with OAA to form citrate and CoA, the first step in the citric acid cycle (TCA or Krebs cycle). The overall CS reaction is thought to proceed through three partial reactions and involves both closed and open conformational forms of the enzyme: a) the carbanion or equivalent is generated from AcCoA by base abstraction of a proton, b) the nucleophilic attack of this carbanion on OAA to generate citryl-CoA, and c) the hydrolysis of citryl-CoA to produce citrate and CoA. There are two types of CSs: type I CS and type II CSs. Type I CSs are found in eukarya, gram-positive bacteria, archaea, and in some gram-negative bacteria and are homodimers with both subunits participating in the active site. Type II CSs are unique to gram-negative bacteria and are homohexamers of identical subunits (approximated as a trimer of dimers). ScCit3 is mitochondrial and functions in the metabolism of PrCoA; it is a dual specificity CS and 2MCS, having similar catalytic efficiency with both AcCoA and PrCoA. The pattern of expression of the ScCIT3 gene follows that of the major mitochondrial CS gene (CIT1, not included in this group) and its expression is increased in the presence of a CIT1 deletion. This group also contains Aspergillus nidulans 2MCS; a deletion of the gene encoding this protein results in a strain unable to grow on propionate. This group contains proteins which functions exclusively as either a CS or a 2MCS, as well as those with relaxed specificity which have dual functions as both a CS and a 2MCS. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.