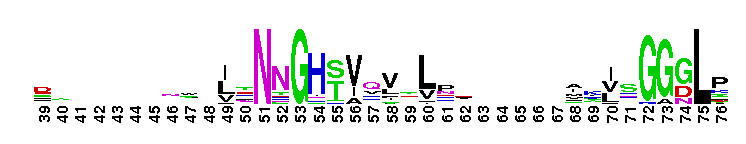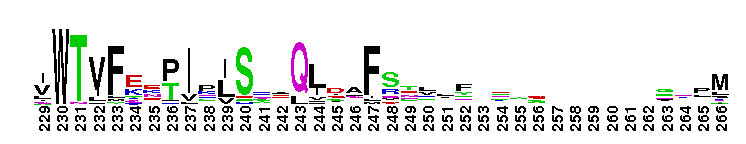| |||||||||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




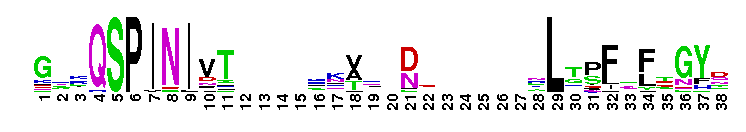
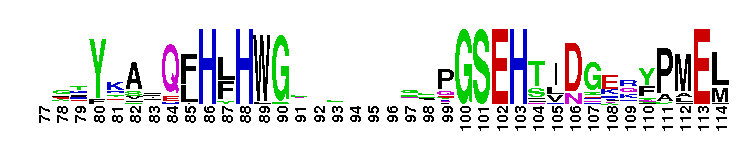
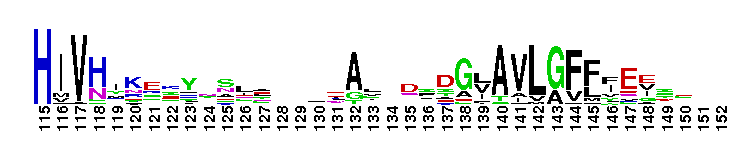
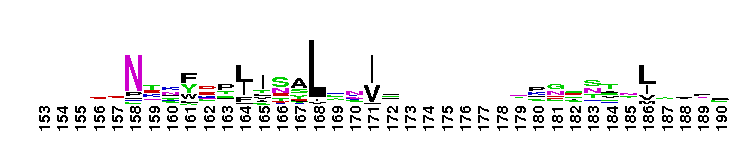
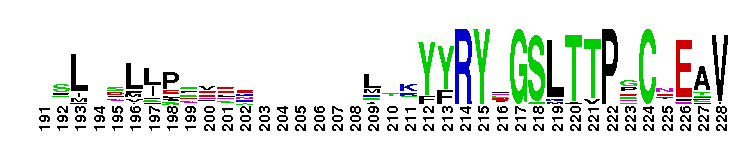
 Carbonic anhydrase alpha, CA_IV, CA_XV, like isozymes. Carbonic anhydrases (CAs) are zinc-containing enzymes that catalyze the reversible hydration of carbon dioxide in a two-step mechanism: a nucleophilic attack of a zinc-bound hydroxide ion on carbon dioxide, followed by the regeneration of the active site by ionization of the zinc-bound water molecule and removal of a proton from the active site. They are ubiquitous enzymes involved in fundamental processes like photosynthesis, respiration, pH homeostasis and ion transport. There are three evolutionary distinct groups - alpha, beta and gamma carbonic anhydrases - which show no significant sequence identity or structural similarity. Most alpha CAs are monomeric enzymes. The zinc ion is complexed by three histidine residues. This subgroup, restricted to animals, contains isozyme IV and similar proteins such as mouse CA XV. Isozymes IV is attached to membranes via a glycosylphosphatidylinositol (GPI) tail. In mammals, Isozyme IV plays crucial roles in kidney and lung function, amongst others. This subgroup also contains the dual domain CA from the giant clam, Tridacna gigas. T. gigas CA plays a role in the movement of inorganic carbon from the surrounding seawater to the symbiotic algae found in the clam's tissues. CA XV is expressed in several species but not in humans or chimps. Similar to isozyme CA IV, CA XV attaches to membranes via a GPI tail.
Carbonic anhydrase alpha, CA_IV, CA_XV, like isozymes. Carbonic anhydrases (CAs) are zinc-containing enzymes that catalyze the reversible hydration of carbon dioxide in a two-step mechanism: a nucleophilic attack of a zinc-bound hydroxide ion on carbon dioxide, followed by the regeneration of the active site by ionization of the zinc-bound water molecule and removal of a proton from the active site. They are ubiquitous enzymes involved in fundamental processes like photosynthesis, respiration, pH homeostasis and ion transport. There are three evolutionary distinct groups - alpha, beta and gamma carbonic anhydrases - which show no significant sequence identity or structural similarity. Most alpha CAs are monomeric enzymes. The zinc ion is complexed by three histidine residues. This subgroup, restricted to animals, contains isozyme IV and similar proteins such as mouse CA XV. Isozymes IV is attached to membranes via a glycosylphosphatidylinositol (GPI) tail. In mammals, Isozyme IV plays crucial roles in kidney and lung function, amongst others. This subgroup also contains the dual domain CA from the giant clam, Tridacna gigas. T. gigas CA plays a role in the movement of inorganic carbon from the surrounding seawater to the symbiotic algae found in the clam's tissues. CA XV is expressed in several species but not in humans or chimps. Similar to isozyme CA IV, CA XV attaches to membranes via a GPI tail. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.