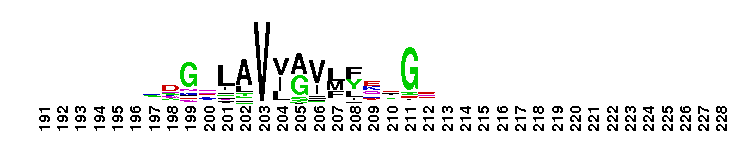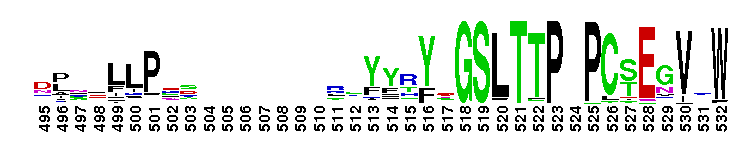| |||||||||||||||||||||||||
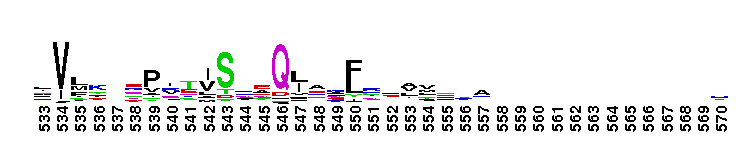
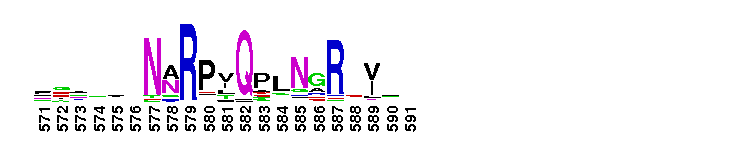
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




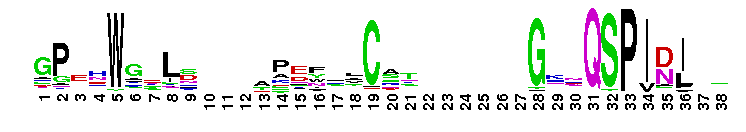
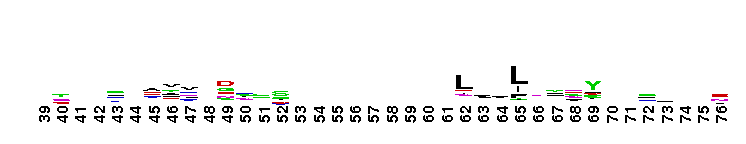
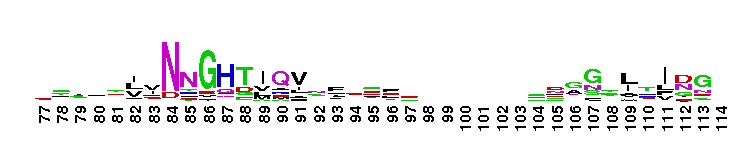
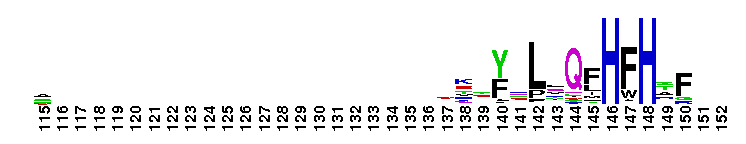
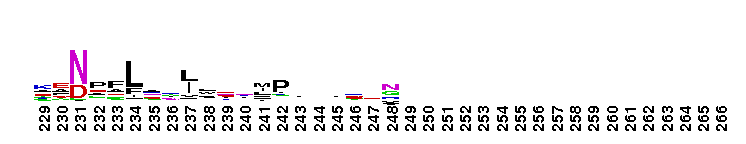
 Carbonic anhydrase alpha, prokaryotic-like subfamily. Carbonic anhydrases (CAs) are zinc-containing enzymes that catalyze the reversible hydration of carbon dioxide in a two-step mechanism: a nucleophilic attack of a zinc-bound hydroxide ion on carbon dioxide, followed by the regeneration of the active site by ionization of the zinc-bound water molecule and removal of a proton from the active site. They are ubiquitous enzymes involved in fundamental processes like photosynthesis, respiration, pH homeostasis and ion transport. Most alpha CAs are monomeric enzymes. The zinc ion is complexed by three histidines. This sub-family includes bacterial carbonic anhydrase alpha, as well as plant enzymes such as tobacco nectarin III and yam dioscorin and, carbonic anhydrases from molluscs, such as nacrein, which are part of the organic matrix layer in shells. Other members of this family may be involved in maintaining pH balance, in facilitating transport of carbon dioxide or carbonic acid, or in sensing carbon dioxide levels in the environment. Dioscorin is the major storage protein of yam tubers and may play a role as an antioxidant. Tobacco Nectarin may play a role in the maintenace of pH and oxidative balance in nectar. Mollusc nacrein may participate in calcium carbonate crystal formation of the nacreous layer. This subfamily also includes three alpha carbonic anhydrases from Chlamydomonas reinhardtii (CAH 1-3). CAHs1-2 are localized in the periplasmic space. CAH1 faciliates the movement of carbon dioxide across the plasma membrane when the medium is alkaline. CAH3 is localized to the thylakoid lumen and provides CO2 to Rubisco.
Carbonic anhydrase alpha, prokaryotic-like subfamily. Carbonic anhydrases (CAs) are zinc-containing enzymes that catalyze the reversible hydration of carbon dioxide in a two-step mechanism: a nucleophilic attack of a zinc-bound hydroxide ion on carbon dioxide, followed by the regeneration of the active site by ionization of the zinc-bound water molecule and removal of a proton from the active site. They are ubiquitous enzymes involved in fundamental processes like photosynthesis, respiration, pH homeostasis and ion transport. Most alpha CAs are monomeric enzymes. The zinc ion is complexed by three histidines. This sub-family includes bacterial carbonic anhydrase alpha, as well as plant enzymes such as tobacco nectarin III and yam dioscorin and, carbonic anhydrases from molluscs, such as nacrein, which are part of the organic matrix layer in shells. Other members of this family may be involved in maintaining pH balance, in facilitating transport of carbon dioxide or carbonic acid, or in sensing carbon dioxide levels in the environment. Dioscorin is the major storage protein of yam tubers and may play a role as an antioxidant. Tobacco Nectarin may play a role in the maintenace of pH and oxidative balance in nectar. Mollusc nacrein may participate in calcium carbonate crystal formation of the nacreous layer. This subfamily also includes three alpha carbonic anhydrases from Chlamydomonas reinhardtii (CAH 1-3). CAHs1-2 are localized in the periplasmic space. CAH1 faciliates the movement of carbon dioxide across the plasma membrane when the medium is alkaline. CAH3 is localized to the thylakoid lumen and provides CO2 to Rubisco. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.