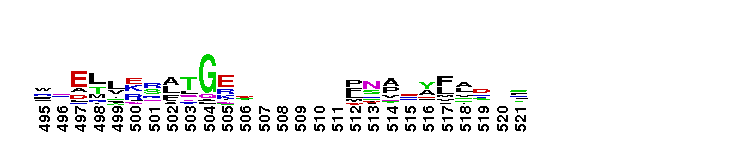| |||||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




 Peptidase M3-like family, a zincin metallopeptidase, includes M3 and M32 families. The peptidase M3-like family, also called neurolysin-like family, is part of the "zincins" metallopeptidases, and includes M3 and M32 families of metallopeptidases. The M3 family is subdivided into two subfamilies: the widespread M3A, which comprises a number of high-molecular mass endo- and exopeptidases from bacteria, archaea, protozoa, fungi, plants and animals, and the small M3B, whose members are enzymes primarily from bacteria. Well-known mammalian/eukaryotic M3A endopeptidases are the thimet oligopeptidase (TOP; endopeptidase 3.4.24.15), neurolysin (alias endopeptidase 3.4.24.16), and the mitochondrial intermediate peptidase. The first two are intracellular oligopeptidases, which act only on relatively short substrates of less than 20 amino acid residues, while the latter cleaves N-terminal octapeptides from proteins during their import into the mitochondria. The M3A subfamily also contains several bacterial endopeptidases, collectively called oligopeptidases A, as well as a large number of bacterial carboxypeptidases, called dipeptidyl peptidases (Dcp; Dcp II; peptidyl dipeptidase; EC 3.4.15.5). The peptidases in the M3 family contain the HEXXH motif that forms the active site in conjunction with a C-terminally-located Glutamic acid (Glu) residue. A single zinc ion is ligated by the side-chains of the two Histidine (His) residues, and the more C-terminal Glu. Most of the peptidases are synthesized without signal peptides or propeptides, and function intracellularly. There are similarities to the thermostable carboxypeptidases from Pyrococcus furiosus carboxypeptidase (PfuCP), and Thermus aquaticus (TaqCP), belonging to peptidase family M32. Little is known about function of this family, including carboxypeptidases Taq and Pfu.
Peptidase M3-like family, a zincin metallopeptidase, includes M3 and M32 families. The peptidase M3-like family, also called neurolysin-like family, is part of the "zincins" metallopeptidases, and includes M3 and M32 families of metallopeptidases. The M3 family is subdivided into two subfamilies: the widespread M3A, which comprises a number of high-molecular mass endo- and exopeptidases from bacteria, archaea, protozoa, fungi, plants and animals, and the small M3B, whose members are enzymes primarily from bacteria. Well-known mammalian/eukaryotic M3A endopeptidases are the thimet oligopeptidase (TOP; endopeptidase 3.4.24.15), neurolysin (alias endopeptidase 3.4.24.16), and the mitochondrial intermediate peptidase. The first two are intracellular oligopeptidases, which act only on relatively short substrates of less than 20 amino acid residues, while the latter cleaves N-terminal octapeptides from proteins during their import into the mitochondria. The M3A subfamily also contains several bacterial endopeptidases, collectively called oligopeptidases A, as well as a large number of bacterial carboxypeptidases, called dipeptidyl peptidases (Dcp; Dcp II; peptidyl dipeptidase; EC 3.4.15.5). The peptidases in the M3 family contain the HEXXH motif that forms the active site in conjunction with a C-terminally-located Glutamic acid (Glu) residue. A single zinc ion is ligated by the side-chains of the two Histidine (His) residues, and the more C-terminal Glu. Most of the peptidases are synthesized without signal peptides or propeptides, and function intracellularly. There are similarities to the thermostable carboxypeptidases from Pyrococcus furiosus carboxypeptidase (PfuCP), and Thermus aquaticus (TaqCP), belonging to peptidase family M32. Little is known about function of this family, including carboxypeptidases Taq and Pfu. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.