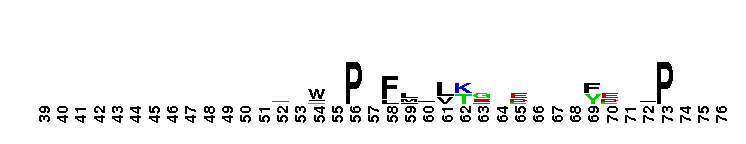| ||||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




 Syntrophin Split Pleckstrin homology (PH) domain. Syntrophins are scaffold proteins that associate with associate with the Duchenne muscular dystrophy protein dystrophin and the dystrophin-related proteins, utrophin and dystrobrevin to form the dystrophin glycoprotein complex (DGC). There are 5 members: alpha, beta1, beta2, gamma1, and gamma2) all of which contains a split (also called joined) PH domain and a PDZ domain (PHN-PDZ-PHC). The split PH domain of alpha-syntrophin adopts a canonical PH domain fold and together with PDZ forms a supramodule functioning synergistically in binding to inositol phospholipids. The alpha-syntrophin PH-PDZ supramodule showed strong binding to phosphoinositides PI(3,5)P2 and PI(5)P, modest binding to PI(3,4)P2 and PI(4,5)P2, and weak binding to PI(3)P, PI(4)P, and PI(3,4,5)P. There are a large number of signaling proteins that bind to the PDZ domain of syntrophins: nitric oxide synthase (nNOS), aquaporin-4, voltage-gated sodium channels, potassium channels, serine/threonine protein kinases, and the ATP-binding cassette transporter A1. PH domains have diverse functions, but in general are involved in targeting proteins to the appropriate cellular location or in the interaction with a binding partner. They share little sequence conservation, but all have a common fold, which is electrostatically polarized. Less than 10% of PH domains bind phosphoinositide phosphates (PIPs) with high affinity and specificity. PH domains are distinguished from other PIP-binding domains by their specific high-affinity binding to PIPs with two vicinal phosphate groups: PtdIns(3,4)P2, PtdIns(4,5)P2 or PtdIns(3,4,5)P3 which results in targeting some PH domain proteins to the plasma membrane. A few display strong specificity in lipid binding. Any specificity is usually determined by loop regions or insertions in the N-terminus of the domain, which are not conserved across all PH domains. PH domains are found in cellular signaling proteins such as serine/threonine kinase, tyrosine kinases, regulators of G-proteins, endocytotic GTPases, adaptors, as well as cytoskeletal associated molecules and in lipid associated enzymes.
Syntrophin Split Pleckstrin homology (PH) domain. Syntrophins are scaffold proteins that associate with associate with the Duchenne muscular dystrophy protein dystrophin and the dystrophin-related proteins, utrophin and dystrobrevin to form the dystrophin glycoprotein complex (DGC). There are 5 members: alpha, beta1, beta2, gamma1, and gamma2) all of which contains a split (also called joined) PH domain and a PDZ domain (PHN-PDZ-PHC). The split PH domain of alpha-syntrophin adopts a canonical PH domain fold and together with PDZ forms a supramodule functioning synergistically in binding to inositol phospholipids. The alpha-syntrophin PH-PDZ supramodule showed strong binding to phosphoinositides PI(3,5)P2 and PI(5)P, modest binding to PI(3,4)P2 and PI(4,5)P2, and weak binding to PI(3)P, PI(4)P, and PI(3,4,5)P. There are a large number of signaling proteins that bind to the PDZ domain of syntrophins: nitric oxide synthase (nNOS), aquaporin-4, voltage-gated sodium channels, potassium channels, serine/threonine protein kinases, and the ATP-binding cassette transporter A1. PH domains have diverse functions, but in general are involved in targeting proteins to the appropriate cellular location or in the interaction with a binding partner. They share little sequence conservation, but all have a common fold, which is electrostatically polarized. Less than 10% of PH domains bind phosphoinositide phosphates (PIPs) with high affinity and specificity. PH domains are distinguished from other PIP-binding domains by their specific high-affinity binding to PIPs with two vicinal phosphate groups: PtdIns(3,4)P2, PtdIns(4,5)P2 or PtdIns(3,4,5)P3 which results in targeting some PH domain proteins to the plasma membrane. A few display strong specificity in lipid binding. Any specificity is usually determined by loop regions or insertions in the N-terminus of the domain, which are not conserved across all PH domains. PH domains are found in cellular signaling proteins such as serine/threonine kinase, tyrosine kinases, regulators of G-proteins, endocytotic GTPases, adaptors, as well as cytoskeletal associated molecules and in lipid associated enzymes. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.