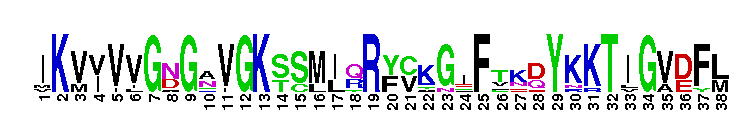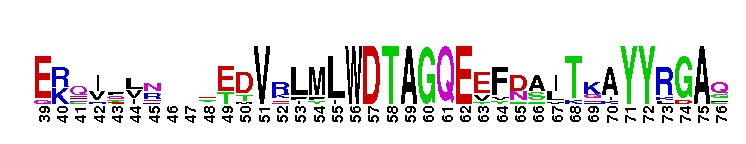| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Tips:  Range on the Protein: Protein ID Protein Position Domain Position: 
|
|---|
Weblogos are Copyright (c) 2002 Regents of the University of California
| DMDM_info@umbc.edu | 1000 Hilltop Circle, Baltimore, MD 21250 | Department of Biological Sciences | Phone: 410-455-2258 |




 Rab GTPase family 23 (Rab23)-like. Rab23-like subfamily. Rab23 is a member of the Rab family of small GTPases. In mouse, Rab23 has been shown to function as a negative regulator in the sonic hedgehog (Shh) signaling pathway. Rab23 mediates the activity of Gli2 and Gli3, transcription factors that regulate Shh signaling in the spinal cord, primarily by preventing Gli2 activation in the absence of Shh ligand. Rab23 also regulates a step in the cytoplasmic signal transduction pathway that mediates the effect of Smoothened (one of two integral membrane proteins that are essential components of the Shh signaling pathway in vertebrates). In humans, Rab23 is expressed in the retina. Mice contain an isoform that shares 93% sequence identity with the human Rab23 and an alternative splicing isoform that is specific to the brain. This isoform causes the murine open brain phenotype, indicating it may have a role in the development of the central nervous system. GTPase activating proteins (GAPs) interact with GTP-bound Rab and accelerate the hydrolysis of GTP to GDP. Guanine nucleotide exchange factors (GEFs) interact with GDP-bound Rabs to promote the formation of the GTP-bound state. Rabs are further regulated by guanine nucleotide dissociation inhibitors (GDIs), which facilitate Rab recycling by masking C-terminal lipid binding and promoting cytosolic localization. Most Rab GTPases contain a lipid modification site at the C-terminus, with sequence motifs CC, CXC, or CCX. Lipid binding is essential for membrane attachment, a key feature of most Rab proteins. Due to the presence of truncated sequences in this CD, the lipid modification site is not available for annotation.
Rab GTPase family 23 (Rab23)-like. Rab23-like subfamily. Rab23 is a member of the Rab family of small GTPases. In mouse, Rab23 has been shown to function as a negative regulator in the sonic hedgehog (Shh) signaling pathway. Rab23 mediates the activity of Gli2 and Gli3, transcription factors that regulate Shh signaling in the spinal cord, primarily by preventing Gli2 activation in the absence of Shh ligand. Rab23 also regulates a step in the cytoplasmic signal transduction pathway that mediates the effect of Smoothened (one of two integral membrane proteins that are essential components of the Shh signaling pathway in vertebrates). In humans, Rab23 is expressed in the retina. Mice contain an isoform that shares 93% sequence identity with the human Rab23 and an alternative splicing isoform that is specific to the brain. This isoform causes the murine open brain phenotype, indicating it may have a role in the development of the central nervous system. GTPase activating proteins (GAPs) interact with GTP-bound Rab and accelerate the hydrolysis of GTP to GDP. Guanine nucleotide exchange factors (GEFs) interact with GDP-bound Rabs to promote the formation of the GTP-bound state. Rabs are further regulated by guanine nucleotide dissociation inhibitors (GDIs), which facilitate Rab recycling by masking C-terminal lipid binding and promoting cytosolic localization. Most Rab GTPases contain a lipid modification site at the C-terminus, with sequence motifs CC, CXC, or CCX. Lipid binding is essential for membrane attachment, a key feature of most Rab proteins. Due to the presence of truncated sequences in this CD, the lipid modification site is not available for annotation. No pairwise interactions are available for this conserved domain.
No pairwise interactions are available for this conserved domain.